Welcome!
Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.

Dr. Ed Kim from the Levine Cancer Institute in Charlotte, NC summarizes the mechanism of next generation sequencing (NGS), how it can potentially be used, and its limitations in clinical practice today.
Transcript
As you know, this is a very, very broad field now — the terminology almost gets mixed into any type of platform testing, or diagnostic testing you’ll hear about, and it’s difficult because it’s not something that’s exclusive, that can be ordered by doctors. This is something that also patients request. There are companies coming from all different directions that can tell you, “this is the test you should order,” or “it could be out of your blood,” “it could be out of your tissue,” and I think it’s important to put it in the context that some of these markers are very useful, but some of them are a little bit more limited. So it’s really good to have a discussion with your physician about what test is appropriate to help determine the best treatment.
Now, if we talk about next-gen sequencing — that’s where the NGS comes from, so any time you hear about NGS, that is what we’re talking about. Now, in the early 2000s and before, what most technologies used was what we call Sanger sequencing, and I really don’t want you to get too tied up into the terminology, but essentially, older technology was like building a railroad — you had to lay down each little piece of the track, one by one, and do it in a sequential manner. You can imagine if you are actually measuring your entire genetic code, that can take a long time — it’s a very exhaustive process, and certainly can miss some things.
With NGS, now, it’s like laying down parallel tracks with railroad producing machines, and so you can lay down multiple tracks at the same time. Now, what you are doing is doing multiple genes in sequence, and in parallel. So, it is a much more efficient method, you’re not missing as many things, and it’s a little less labor intensive — and that’s what has really changed why we can get a lot of genes, or information from your genes, in a quicker, efficient manner, because of this type of platform technology. The other name of it was massively parallel sequencing, but that doesn’t sound as sexy as NGS, so NGS is the terminology that will help you identify what next-gen sequencing and that type of testing is.
There are multiple names of platforms that exist out there, that go under this NGS, or conducting next-gen sequencing: I mentioned Sanger, there’s also Illumina MiSeq, there’s also a Roche 454 sequencing, you also hear terms like ion torrent and ion proton — that also does sequencing, and also solid sequencing. So, just to throw out some names, just in case you hear some of these, just know that they all essentially try to accomplish the same things; some will have more genes, some will have less genes, but what they’re trying to do is identify what abnormalities may exist within the tumor’s DNA, or genes.
As researchers, and scientists, and clinicians, we really like having this type of technology. It really allows us to take a tumor sample from a patient, and study it, measuring thousands and thousands of different genes. However, we know that there are limitations, and we can’t measure all of them — it is very time consuming, and there is cost associated with it. So we try to embed these in the context of clinical research as much as we can, in order to really learn how a tumor behaves, how it reacts to certain therapies, and how it affects the patient overall. It is a useful tool, and you’ll see it and hear about it more and more.
Please feel free to offer comments and raise questions in our
discussion forums.
Hi app.92, Welcome to Grace. I'm sorry this is late getting to you. And more sorry your mum is going through this. It's possible this isn't a pancoast tumor even though...
A Brief Tornado. I love the analogy Dr. Antonoff gave us to describe her presentation. I felt it earlier too and am looking forward to going back for deeper dive.
Dr. Singhi's reprise on appropriate treatment, "Right patient, right time, right team".
While Dr. Ryckman described radiation oncology as "the perfect blend of nerd skills and empathy".
I hope any...
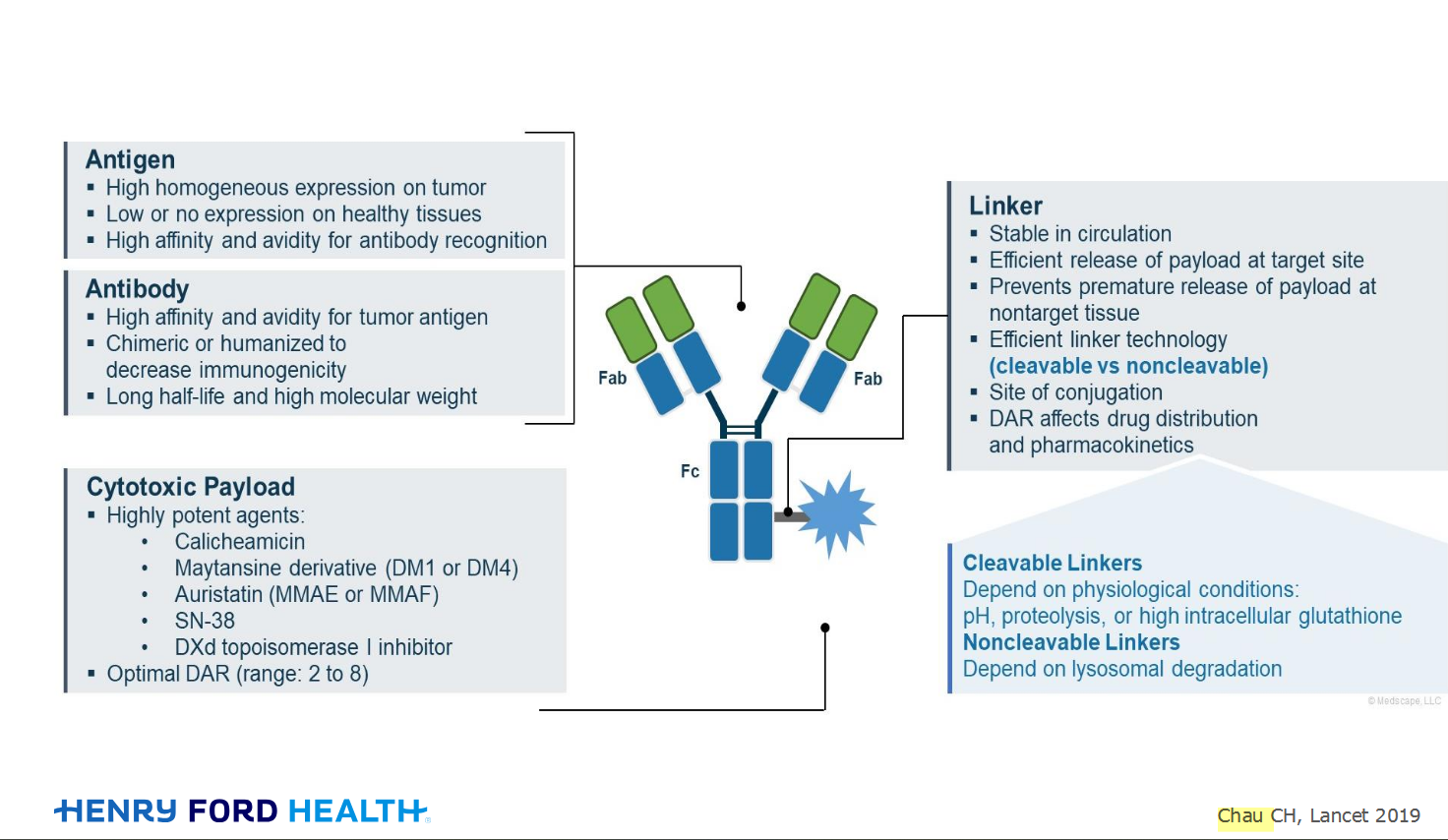
My understanding of ADCs is very basic. I plan to study Dr. Rous’ discussion to broaden that understanding.
Here's the webinar on YouTube. It begins with the agenda. Note the link is a playlist, which will be populated with shorts from the webinar on specific topics
An antibody–drug conjugate (ADC) works a bit like a Trojan horse. It has three main components:

Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.