Welcome!
Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.

Drs. Leora Horn, Ben Solomon, & Jack West discuss the open question of whether there are clinically significant differences among leading EGFR tyrosine kinases based on the specific EGFR mutation to be treated.
[powerpress]
[ratingwidget post_id=0]
Please feel free to offer comments and raise questions in our Discussion Forums.
Transcript
Dr. West: What about rare mutations? They’ve really been excluded from a lot of things, but we got a little bit of information at this meeting, the World Conference on Lung Cancer, and there was a publication by James Yang in Lancet Oncology that looked at aggregate results from three different studies with afatinib (Gilotrif), that suggested that at least some of these rare mutations could have a good response to afatinib. What are your thoughts — do you really view all of these rare mutations as one basket, or do you hone it further and treat accordingly, and if you do feel that there are patients who benefit from EGFR inhibitors, is it preferentially afatinib because of these results, or any of them? Leora, what do you think?
Dr. Horn: So, I thought that James Yang paper was very nice, but I think my bias is that, probably, those rare mutations, would have responded to the first generation inhibitors, as well. We have our database that we’re collecting this data — the DIRECT database, and we’ve got about 3,000 individual mutation patient information in there, so that’s my go-to place when I get information that I’m not sure what to make of a mutation, and I think we’re going to see more and more rare mutations as next generation sequencing is adopted in more places around the world, and we may not always know what to do with that information, and sometimes it may be trial and error.
Dr. Solomon: Yeah, so I actually have to say your mycancergenome.org is my go-to place, as well, to find out about rare mutations, and I think for patients, and even doctors, who get an EGFR mutation result with a rare mutation, it’s hard to keep track of all these mutations, and at this meeting, for instance, we learned that the group of mutations we call G719X, which includes three different mutations — two of those mutations seem to respond, and one of those, the G719C, doesn’t seem to respond, and so I would encourage people who are uncertain about the significance of mutations to go to mycancergenome.org and make query of the DIRECT database.
Dr. West: To me, that was one of the themes, both from the Yang paper, and from several of the still relatively small data presentations on rare mutations, is that, de novo T790m mutations, that is, the ones that appear from the time of diagnosis, are associated with a poor outcome, but some of the others, Exon 18, and some Exon 20 insertions, could respond well to these therapies, and it would be a mistake to: A., lump all of these together, because they’re not, we’re missing things, and B., to completely discount the possibility that an EFGR inhibitor could help them.
Please feel free to offer comments and raise questions in our
discussion forums.
Dr. Singhi's reprise on appropriate treatment, "Right patient, right time, right team".
While Dr. Ryckman described radiation oncology as "the perfect blend of nerd skills and empathy".
I hope any...
My understanding of ADCs is very basic. I plan to study Dr. Rous’ discussion to broaden that understanding.
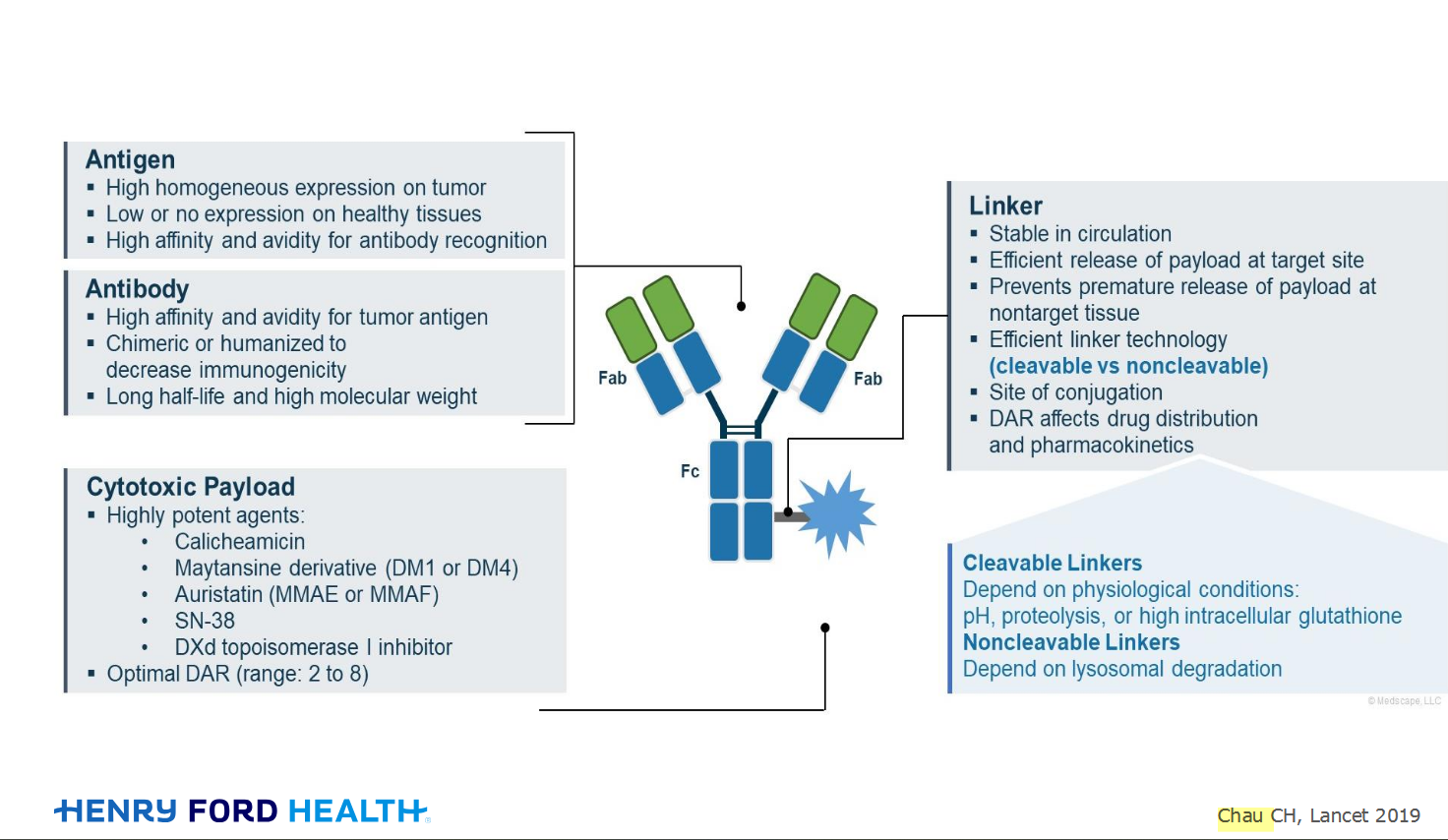
An antibody–drug conjugate (ADC) works a bit like a Trojan horse. It has three main components:
Bispecifics, or bispecific antibodies, are advanced immunotherapy drugs engineered to have two binding sites, allowing them to latch onto two different targets simultaneously, like a cancer cell and a T-cell, effectively...
The prefix “oligo–” means few. Oligometastatic (at diagnosis) Oligoprogression (during treatment)
There will be a discussion, “Studies in Oligometastatic NSCLC: Current Data and Definitions,” which will focus on what we...
Radiation therapy is primarily a localized treatment, meaning it precisely targets a specific tumor or area of the body, unlike systemic treatments (like chemotherapy) that affect the whole body.
The...

Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.
A Brief Tornado. I love the analogy Dr. Antonoff gave us to describe her presentation. I felt it earlier too and am looking forward to going back for deeper dive.