Welcome!
Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.
Continuing with the analysis of a publication about tarceva (erlotinib) for patients with advanced BAC that I introduced in the last post, we'll turn now to the analysis that Dr. Vince Miller and colleagues did on the biomarkers that might predict more or less clinical benefit with an EGFR inhibitor like tarceva (abstract here). The trial looked at three different ways of measuring EGFR:
1) EGFR activating mutations in the gene, which are located in the part of the EGFR molecule where drugs like tarceva and iressa (gefitinib) work, and which have been associated with a high likelihood of response to these agents
2) EGFR "copy number", which measures overall amplification of the number of copies of the gene in cancer cells, and which at high levels has been shown by some groups to be associated with a higher likelihood of response and prolonged survival than low/normal levels; this was measured here by "chromogenic in situ hybridization, or CISH, similar to the more commonly used test called fluorescence in situ hybridization (FISH)
3) EGFR immunohistochemistry (IHC), which measures the amount of the protein on the surface of the cancer cells, and which has been least clearly associated with better or worse results with EGFR inhibitors, at least the oral tyrosine kinase inhibitors like tarceva and iressa.
In addition, the investigators also looked at K-RAS mutations in the cancer, which have been associated with smoking and rarely seen in never-smokers, and they've also been associated with resistance to EGFR inhibitors and likely also many types of chemotherapy. K-RAS mutations appear to be associated with resistance and poor response in general, and some people have suggested that EGFR inhibitors not be tried in patients with K-RAS mutations, because responses have rarely if ever been seen in patients with K-RAS mutation-positive cancers.
Of the 101 patients with advanced BAC and treated with tarceva on the trial, 82 had enough tissue for at least one of these biomarker studies, and 61 had enough tissue collected to perform all four studies. As with several prior studies that have looked at EGFR mutations, the 22% of patients with EGFR activating mutations (higher than expected for a North American population, probably because EGFR mutations are more common in BAC than in lung cancer in general) had a response rate (RR) of 83%, compared with only 7% for patients without EGFR mutations ("wild type"). This highly statistically significant difference was also mirrored by a significantly longer median progression-free survival (PFS) among EGFR mutation-positive patients (13 vs. 2 months), but the difference became attentuated and wasn't statistically signficant for median overall survival (OS) (23 vs. 17 months). Gene amplification by CISH showed the same pattern, but not as dramatically: those patients with EGFR gene amplification by CISH showed a significantly higher RR (43% vs. 13%) and median PFS (9 vs 2 months), but no significant difference in median OS (25 vs. 16 months). And EGFR IHC really didn't show any trends at all for any of these efficacy parameters.
Meanwhile, K-RAS mutations showed what they have tended to: a significant difference in RR, with responses never seen in patients with a tumor harboring a K-RAS mutation (0% vs. 32%)), but no significant differences in median PFS (4 vs. 5 months) or median OS (13 vs. 21 months). As you can see from some of the differences in the numbers, however, the small numbers of patients being compared kept some pretty big differences from being what would be considered "statistically significant" (the larger the trial and the number of people available for comparison, the smaller the differences in outcomes need to be in order to be considered overwhelmingly unlikely to be caused by chance alone).
Another interesting way of looking at response is with a "waterfall plot", in which measured changes in the volume of the cancer with treatment are plotted from largest progression on the left to greatest shrinkage on the right, making what looks like a waterfall. The bars that go downward represent a response (anything from minor to major), and the upward bars are progression:
(Click on image to enlarge)
There are several interesting points here. First, most of the best responses were seen in patients with EGFR mutations, but not all. Second, many of the patients with EGFR mutations also were positive for EGFR by CISH. Third, even though there were no responses among patients with K-RAS mutations, there were patients with these mutations who demonstrated stable disease. Finally, at least half and I would say closer to two thirds of patinets experienced no change or some degree of shrinkage on tarceva, even though the response rate overall was 22%. And we know that survival can be improved in populations with stable disease.
Another interesting issue, though it isn't addressed here, is that we've seen that survival is best for patients with EGFR mutations, apparently no matter what treatment they get. So though we would think that a drug like tarceva or iressa would be the critical driver for improved survival in patients with tumors that have EGFR mutations, they actually seem to just do better overall, so it may be a marker of a generally more responsive and/or slower-growing cancer. The opposite is seen with tumors that have a K-RAS mutation (and we have almost never found a tumor with both an EGFR mutation and a K-RAS mutation): they are not only resistant to EGFR inhibitors, but they also appear to be resistant to conventional chemo as well and just appear to do worse overall.
Although I think the biomarker story is interesting, I would argue that the results here show that you don't need to have an EGFR mutation to respond, that survival is not significantly better for EGFR mutation patients or any other biomarker, and that patients with tumors that show K-RAS mutations may not respond but may show stable disease on tarceva. Taken together, I think the biomarker story is interesting but wouldn't really guide my clinical decisions at this point. I wouldn't only offer tarceva to patients with an EGFR mutation or gene amplification, and I wouldn't necessarily exclude a patient with a K-RAS mutation from getting tarceva either.
As always, I welcome your thoughts and questions.
Please feel free to offer comments and raise questions in our
discussion forums.
A Brief Tornado. I love the analogy Dr. Antonoff gave us to describe her presentation. I felt it earlier too and am looking forward to going back for deeper dive.
Dr. Singhi's reprise on appropriate treatment, "Right patient, right time, right team".
While Dr. Ryckman described radiation oncology as "the perfect blend of nerd skills and empathy".
I hope any...
My understanding of ADCs is very basic. I plan to study Dr. Rous’ discussion to broaden that understanding.
Here's the webinar on YouTube. It begins with the agenda. Note the link is a playlist, which will be populated with shorts from the webinar on specific topics
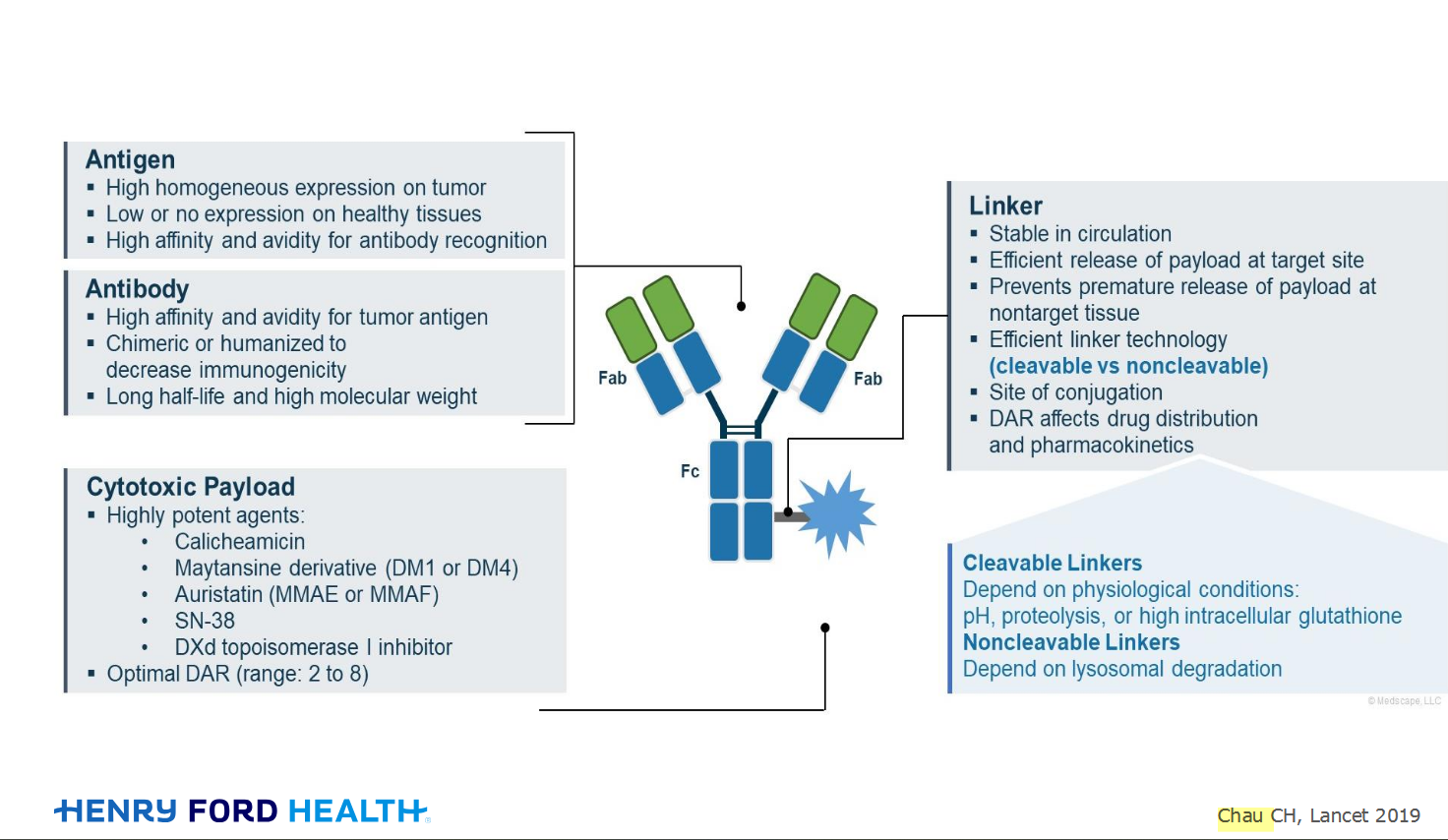
An antibody–drug conjugate (ADC) works a bit like a Trojan horse. It has three main components:
Bispecifics, or bispecific antibodies, are advanced immunotherapy drugs engineered to have two binding sites, allowing them to latch onto two different targets simultaneously, like a cancer cell and a T-cell, effectively...

Welcome to the new CancerGRACE.org! Explore our fresh look and improved features—take a quick tour to see what’s new.
Hi app.92, Welcome to Grace. I'm sorry this is late getting to you. And more sorry your mum is going through this. It's possible this isn't a pancoast tumor even though...